Improving knowledge about metabolites produced by the microbiota is a key point to understand its role in human health and disease. Among them, lipoamino acid (LpAA) containing asparagine and their derivatives are bacterial metabolites which could have an impact on the host.
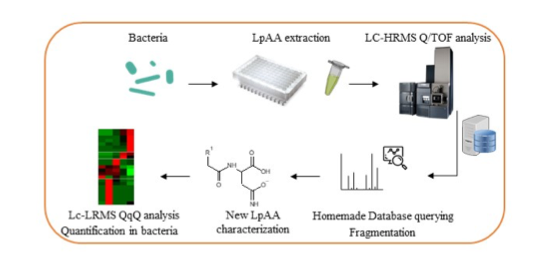
In this study, our aim was to extend the characterization of this family. On MetaToul-Lipidomic faciliy we developed a semi-targeted workflow (from sample preparation to structural characterization by liquid chromatography (LC) coupled to high resolution mass spectrometry (HRMS)) to identify and quantify new candidates. This strategy allowed us to find 25 new LpAA conjugated to Asn, Gln, Asp, Glu, His, Leu, Ileu, Pro, Lys, Phe, Trp and Val amino acids.
These metabolites were then fully characterized by MS2, and compared to the pure synthesized standards to validate annotation. Finally, a quantitative method was developed by LC coupled to a triple quadrupole instrument.
All details can be found in the publication : Hueber, ACA, 2021(doi.org/10.1016/j.aca.2021.339316)


